An open question in gene regulation is how spatio-temporal patterns of gene expression is encoded in the genome. We discussed this paper in our lab meeting. This paper talks about a two-step approach to predicting spatial and temporal gene expression patterns in Drosophila. This prediction task is tackled as a two-step approach: (a) first find cis-regulatory modules that exhibit spatio or temporal activity, (b) second link the crms to predict spatio-temporal expression. Assume space and time is captured by the term “context”. To do (a) the authors use a Bayesian network which is trained on known CRM-context relationships. To do (b) the authors use additional data based on insulators and H3K4me1 to predict which CRMs are associated with which genes. In (a) the CRM-context information is known only for a few hundred of the total 8000 crms that are present. So the authors use an EM idea where they use the trained Bayesian network to predict the context-specific activity of each CRM. Then using the soft labels of each CRM they predict the expression of the gene. This second model is also a Bayesian network but has additional variables for the distance between CRMs and genes and whether there is an insulator binding site.
Category Archives: Chromatin
Chromatin marks and the cell-type specificity of SNPs
This paper came out recently in Nature and combines data from ENCODE, Epigenome consortia, multiple GWAS studies and the 1000 genomes project to address the question of cell type specificity of genetic variation affecting diseases.
The authors try to get to two related questions: (a) what cell types are associated with a diseasem, by looking at the chromatin activities surrounding SNPs associated with a disease, (b) what marks are conferring this cell-type specificity to a disease, and such marks are called the informative marks. It all boils down to computing a statistic that measures how variable the strength of a mark is for SNPs in a disease.
The authors started off with SNPS associated with different diseases from a GWAS study. This analysis was done in a per-disease basis, for example consider LDL or rheumatoid arthritis, etc. The authors found what SNPS are associated with these studies in a GWAS study and added to this list some more SNPs that were in high linkage disequilibrium with these associated SNPs. Then they obtained chromatin mark peaks for different chromatin parks in different cell types and lines from ENCODE as well as the epigenome map. Then they asked for each SNP to what extent were they associated with a particular mark in a particular cell type. This was done by defining a score which is the ratio of the height of a peak to the width of the peak.
Thus if we were to think of this data as a matrix, we would have one matrix per mark, whose columns correspond to the positions of the SNPs and the rows correspond to differnce cell types. A mark is then considered informative for a disease and cell type if all or most of the marks exhibit a high score for a few cell types. A mark is uninformative if the snps associated with the highest scores are not the same across different cell types. To compute this score of informativeness of a mark, the authors defined a metric which measures the variation in the score of SNPs for a disease across cell types. Specifically, the statistic is a sum of square differences of SNP score, and the differences are computed for each cell type and phenotype combination. If this number is small, then the mark is apparently cell-type specific. Finally the authors use a pemutation analysis to identify whether a particular score is high or low. cell-type specificity for a disease is computed by summing over the scores over all snps in a given cell type and assessing significance.
The statistic they use to define whether a mark is informative is the sum of squared differences between each snp’s score and the mean of all snps in each disease cell type combination. If this is small, then we can assume that the mark does not vary too much, but there is no control over which cell types the mark must vary over. I am not sure how the method deals with the situation where a mark is not changing a lot across cell types.
Directionality and heterogeneity of regulatory elements
A new paper from Kundaje et al., released as part of ENCODE, analyzes the arrangement of nucleosomes and histone modifications around transcription start sites (TSS) and transcription factor binding sites (TFBS) in two human tissues. Their study is motivated by the standard aggregation plot, which aggregates genome-wide profiles of a particular signal (say, MNase-seq measures of nucleosome occupancy) around a ubiquitous anchor, such as TSS’s. These aggregation plots show the general features of signal behavior around the anchor (in this case, the customary nucleosome peaks before and after the TSS), but fail to describe the diversity of signal patterns that contribute to this aggregation. Additionally, it aims to describe anchor points where the directionality is known (such as TSS, which are oriented in the direction of transcription) as well as unknown (such as distal TFBS like CTCF).
They create a new tool, Clustered AGgregation Tool (CAGT), which automatically detects distinct clusters of nucleosome or histone modification signal around a regulatory element. CAGT has two major components. First, k-medians clustering is applied to a region of a given size around each anchor point to produce a large set of signal patterns. K-medians requires only a distance metric and a choice of k; they use one minus the base-wise correlation between two signals to quantify their distance. An example cluster might contain a strong peak a set distance to the left of the anchor, and no peak on the right. K-medians might produce a large number of redundant clusters, and does not appropriately flip and combine clusters centered on anchors with unknown polarity. Therefore, the second step of CAGT is hierarchical clustering with the option of reversing signals when the anchors have unknown direction. This produces a consensus set of distinct, diverse nucleosome or chromatin mark signals around anchors such as TSS or TFBS.
They apply their method to human GM12878 and K562 cells. They find diverse positioning patterns for nucleosomes as well as various chromatin marks around gene TSS as well as numerous TFBS sites. The majority of these patterns are assymetrical, suggesting that the inherent polarity of regulatory elements strongly influences the position of these signals. Exceptions included DNase hypersensitivity signals, which were generally symmetric around all TFBS. Interestingly, nucleosome positions anchored around CTCF/cohesin complex binding sites were the most symmetrical of all the TFBS measured, suggesting a unique chromatin environment around insulator elements. In order to perform their computational analysis, they produce high-quality nucleosome positioning data genome-wide for GM12878 and K562 cells using MNase-seq.
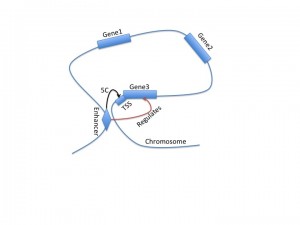
Using 5C to investigate TSS-enhancer interactions
A recent paper from the Dekker lab, released as part of the ENCODE project, describes 5C, a new tool for analyzing three-dimensional looping interactions in chromosomes at unprecedented resolution. 5C, which stands for chromosome conformation capture carbon copy, is capable of describing interactions between promoter regions and distal regulatory regions, providing a new clue for connecting regulatory regions to the genes that they regulate. This technology is still limited by the number of experiments required to investigate large regions of the genome; in this study, they examined only 1% of the genome, corresponding to the ENCODE pilot regions.
This study is motivated by the difficulty in assigning regulatory regions to target genes. They used genome annotations from another ENCODE paper to divide the genome into enhancers, promoters, CTCF, and other sections, and investigated the three-dimensional relationships between TSS and these regions. Unlike promoters, distal enhancers do not necessarily correspond to the nearest gene. This paper finds that only 7% of looping interactions are between an enhancer and the nearest TSS, and 22% of looping interactions are between an enhancer and the nearest active TSS. This supports the idea that genes within the ‘loop’ section of the chromosome structure may not be regulated by the enhancer that regulates a gene at the ‘narrow’ part of the loop.
Interestingly, they found that enhancers between enhancer, promoter, and CTCF regions were most common about 120kb upstream of the TSS for a gene. Less surprisingly, they found that TSS with more 5C interactions are more highly expressed. Furthermore, the 3d interaction network is tissue-specific, although this is more true for TSS-promoter and TSS-enhancer interactions than TSS-CTCF. 5C is still far from perfect – in particular, we still do not have a 5C map of the entire genome. However, this data helps fill in the gaps from more traditional 3C, 4C, and Hi-C experiments, and provides novel insight into the role of distal enhancers in gene regulation.
Segway – a tool for unsupervised segmentation of the human genome
Segway provides a method for automatically segmenting the genome into functional regions by analyzing different kinds of high-throughput data from different experiments. The approach is described in a recent paper from the Noble research lab. Segway uses a Dynamic Bayesian network (DBN) to model the interdependencies between different genomic sections, which is trained using ChIP-seq, DNase-seq, and FAIRE-seq data from ENCODE. They condensed the many discovered segment types into 25 labels which were then assigned functional categories, including familiar terms like gene start, gene middle, gene end, and enhancer. Using this labeling, they recovered many well-known genomic features.
They next compared their results to genome annotations from ChromHMM. While both models produce the same sort of output, the input is different; ChromHMM is trained only with histone modification data, while Segway uses a variety of data types. The authors find that Segway better identifies known elements, has higher segment resolution, and handles missing data better. They focus less on differences across cell type then in the ChromHMM analysis, although their model does appear to accomodate these differences. They conclude by suggesting a hierarchical segmentation approach that could make genome annotation more comprehensible.
ChromHMM – a tool for chromatin state segmentation of genomes
Chromatin marks are an important factor in the transcription regulatory network. A recent study from Ernst et al. uses chip-seq to profile nine distinct histone modifications across nine different human cell types. They developed a tool, ChromHMM, with which they segment the human genome according to the combinatorial pattern of chromatin marks in each segment. ChromHMM applies a multivariate hidden Markov model which models combinations of histone modification using Bernoulli random variables in order to learn a set of distinct chromatin states, and assign each portion of the genome to one state. For their human data, they annotate 15 chromatin states which fall into the broad categories of promoters, enhancers, insulators, transcribed, repressed, and inactive.
They found that enhancer and promoter regions vary widely in activity level between cell types, but that the general categorization of a region as an area of regulatory potential is consistant across tissues. They clustered promoters and enhancers based on chromatin state profile, and found that clusters of promoters are general across cell types, while clusters of enhancers are cell-type specific. Next, they found a strong correlation between patterns of enhancer activity and gene expression levels of the nearest gene, suggesting that distal enhancers often neighbor their target gene. They mapped enhancer regions to target genes based on correlations between enhancer activity profiles, gene expression, sequence motif enrichment, and TF expression. Finally, they found that disease-associated SNPs are significantly enriched in portions of the genome associated with strong enhancer chromatin states.
More information about this study, including the ChromHMM software, can be found on the MIT Computational Biology website.