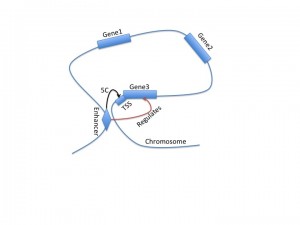
A recent paper from the Dekker lab, released as part of the ENCODE project, describes 5C, a new tool for analyzing three-dimensional looping interactions in chromosomes at unprecedented resolution. 5C, which stands for chromosome conformation capture carbon copy, is capable of describing interactions between promoter regions and distal regulatory regions, providing a new clue for connecting regulatory regions to the genes that they regulate. This technology is still limited by the number of experiments required to investigate large regions of the genome; in this study, they examined only 1% of the genome, corresponding to the ENCODE pilot regions.
This study is motivated by the difficulty in assigning regulatory regions to target genes. They used genome annotations from another ENCODE paper to divide the genome into enhancers, promoters, CTCF, and other sections, and investigated the three-dimensional relationships between TSS and these regions. Unlike promoters, distal enhancers do not necessarily correspond to the nearest gene. This paper finds that only 7% of looping interactions are between an enhancer and the nearest TSS, and 22% of looping interactions are between an enhancer and the nearest active TSS. This supports the idea that genes within the ‘loop’ section of the chromosome structure may not be regulated by the enhancer that regulates a gene at the ‘narrow’ part of the loop.
Interestingly, they found that enhancers between enhancer, promoter, and CTCF regions were most common about 120kb upstream of the TSS for a gene. Less surprisingly, they found that TSS with more 5C interactions are more highly expressed. Furthermore, the 3d interaction network is tissue-specific, although this is more true for TSS-promoter and TSS-enhancer interactions than TSS-CTCF. 5C is still far from perfect – in particular, we still do not have a 5C map of the entire genome. However, this data helps fill in the gaps from more traditional 3C, 4C, and Hi-C experiments, and provides novel insight into the role of distal enhancers in gene regulation.